
Our Research
To understand how Gram-negative bacteria build their outer membranes and protect themselves against antibiotics
The Gram-negative bacterial cell envelope and antimicrobial resistance
Antimicrobial resistant (AMR) micro-organisms cause a major global threat to modern medicine, resulting in over 700,000 deaths annually. Gram-negative, double-membraned bacteria are particularly difficult to treat, in part because their outer membrane (OM) acts as a barrier against antibiotics. Therefore, it is critical to understand how bacteria build and maintain their OM, potentially allowing for novel therapeutic approaches that disrupt this barrier.
The Gram-negative bacterial cell envelope consists of an inner membrane (IM) and an OM separated by an aqueous periplasmic space. To build and maintain this OM, bacteria must transport lipids and hydrophobic proteins across the aqueous periplasm. Increasingly, evidence suggests that membrane components are transported via protein pathways that form shuttles or direct bridges between the IM and OM. We use a combination of structural biology, biochemistry and bacterial genetics to gain important insights into how bacteria build the OM.
The bacterial cell envelope consists of an outer membrane (OM) and an inner membrane (IM), separated by the periplasmic space, which houses the peptidoglycan cell wall. The OM itself is an asymmetric lipid bilayer with an outer leaflet of lipopolysaccharide (LPS) and an inner leaflet of phospholipid (PL). The cell envelope is scattered with proteins that are required for a range of important functions for the bacterial cell, including pathways involved transport of lipids and proteins to build and maintain the OM. Examples depicted are LetB and YhdP, which form tunnels and bridges across the periplasm, respectively.
Recent projects
-
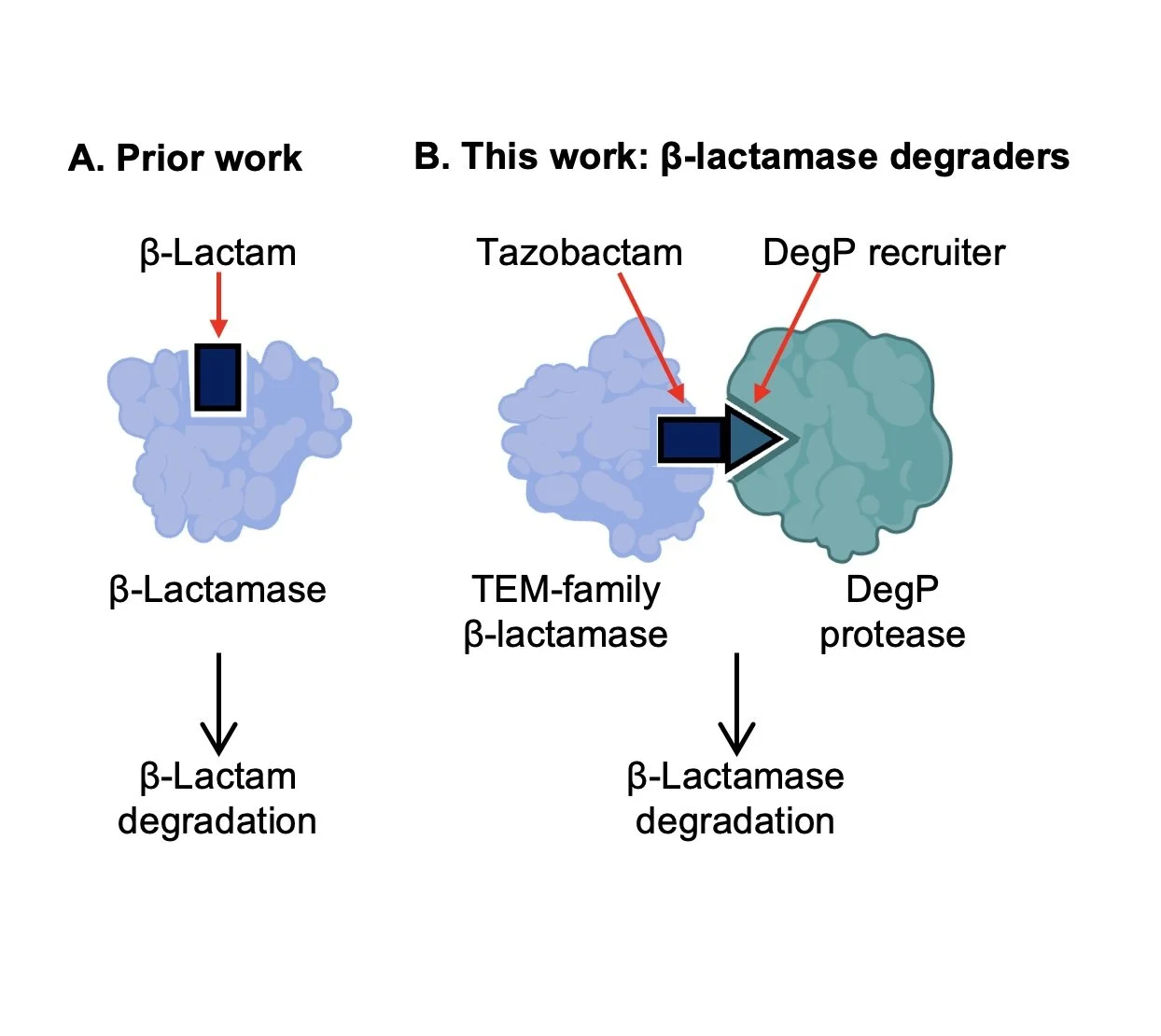
Molecular Glue-like Degraders of β-Lactamases by DegP
Antimicrobial resistance is one of the most serious challenges to global health, yet the development of new molecules with novel mechanisms of action to combat resistance is lacking. Here, we report the discovery of molecular glue-like compounds that recruit TEM-family β-lactamases to the bacterial protease DegP for degradation. β-lactamase inhibitor tazobactam was found to accelerate degradation of TEM β-lactamases by DegP, which was further enhanced by linkerless incorporation of dipeptide motifs enriched among DegP substrates. The resulting molecular glue-like degraders showed improved synergy with β-lactam piperacillin against resistant E. coli compared to tazobactam, as well as good pharmacokinetic properties for oral dosing. Collectively, this work establishes periplasmic targeted protein degradation as a promising new mechanism for combating β-lactamase resistance.
-
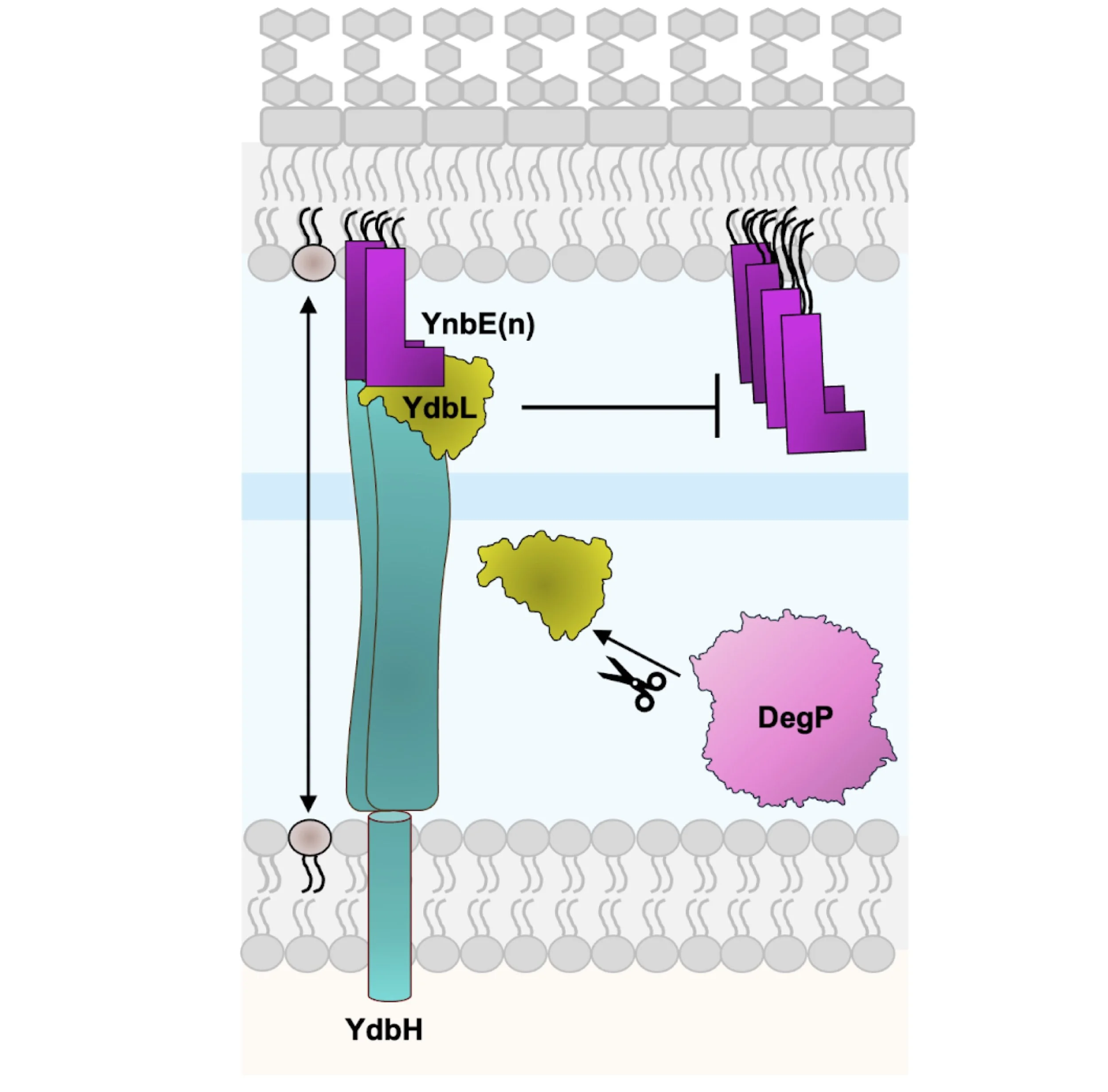
Modulation of the YdbH-YnbE bridge by YdbL
Although the mechanism of phospholipid transport to build the OM remains poorly understood, recent studies implicate TamB, YhdP, and YdbH as functionally redundant and collectively essential proteins mediating this process in Escherichia coli. YdbH is anchored to the inner membrane (IM) and its periplasmic β-sheet groove interacts directly with OM lipoprotein YnbE to form an intermembrane bridge. Our previous data supports a model where YnbE multimerizes to stack on the C-terminus of YdbH, and this multimerization is modulated by the periplasmic protein YdbL. Here, we demonstrate that excess YdbL specifically inhibits the function of the YdbH-YnbE complex. We resolve high-resolution structural data for YdbL and ascertain highly conserved interacting residues with the C-terminus of YnbE, which are critical for YdbL function. Finally, we show that YdbL is protected from degradation by the protease DegP when complexed with YnbE. Overall, our data supports a model in which YdbL directly interacts with YnbE to ensure proper YdbH-YnbE intermembrane bridge formation for lipid transport.
-

YhdP phospholipid transporter
YhdP has been implicated as a major protagonist in the transport of phospholipids to the outer membrane. Here, utilising AlphaFold, we observe YhdP to form an elongated assembly of 60 β strands that curve to form a continuous hydrophobic groove. This architecture is consistent with our negative stain electron microscopy data which reveals YhdP to be approximately 250 Å in length and thus sufficient to span the bacterial cell envelope. Furthermore, molecular dynamics simulations and in vivo bacterial growth assays indicate essential helical regions at the N- and C-termini of YhdP, that may embed into the inner and outer membranes respectively, reinforcing its envelope spanning nature. Our in vivo crosslinking data reveal phosphate-containing substrates captured along the length of the YhdP groove, providing direct evidence that YhdP transports phospholipids. This finding is congruent with our molecular dynamics simulations which demonstrate the propensity for inner membrane lipids to spontaneously enter the groove of YhdP. Collectively, our results support a model in which YhdP bridges the cell envelope, providing a hydrophobic environment for the transport of phospholipids to the outer membrane.
-

MlaFEDB phospholipid transporter
We reveal the cryo-EM structure of lipid transporter MlaFEDB at 3.05 Å resolution. A continuous transport pathway extends from within MlaE, through the channel of MlaD, and into the periplasm. Two phospholipids are bound to MlaFEDB, suggesting that multiple lipid substrates may be transported each cycle. We show that phospholipids can indeed be detected at these locations within the bacterial cell, using an in vivo crosslinking method that we developed.
Read the full story here: N Coudray*, GL Isom*, MR MacRae*, MN Saiduddin, G Bhabha and DC Ekiert (2020). Structure of bacterial phospholipid transporter MlaFEDB with substrate bound. eLife
-

LetB - Lipophilic Envelope Spanning Tunnel
This ~3.5 Å cryo-EM structure of the E. coli MCE protein LetB reveals an ~0.6 megadalton complex that consists of seven stacked rings, with a central hydrophobic tunnel sufficiently long to span the periplasm. Lipids bind inside the tunnel, suggesting that it functions as a pathway for lipid transport. Cryo-EM structures in the open and closed states reveal a dynamic tunnel lining, with implications for gating or substrate translocation.
Read the full story here: GL Isom*, N Coudray*, MR MacRae, CT McManus, DC Ekiert, G Bhabha (2020). LetB structure reveals a tunnel for lipid transport across the bacterial envelope. Cell
Publications
*Co-first author, #Corresponding/last author
C De’Ath*, S Kumar*, S Chhibber, BF Cooper, EK Taylor, E Jones, T Lanyon-Hogg, JR Bolla, C Redfield, N Ruiz# & GL Isom#. YdbL directly modulates YdbH-YnbE bridge formation to maintain Escherichia coli outer membrane homeostasis. BioRxiv
EK Taylor, PS Barbosa, T Kadambi, F Eisele, GL Isom# & Thomas Lanyon-Hogg#. Molecular Glue-like Degraders of TEM β-Lactamases by the Periplasmic Protease DegP. BioRxiv
TCY Seow, JD Bradbury, EK Taylor, LG Bauer, M Harvey, H Hreidarsdottir, SM Saghbini, Y Wang, AM. Thomas, P Santos Barbosa, KVM Huber, GL Isom, T Lanyon-Hogg. Live-Cell Chemoproteomic Profiling Identifies the Uncharacterised Protein YbaA as a Direct Target of Ciprofloxacin in Escherichia coli. BioRxiv
BF Cooper# and GL Isom# (2026). Optimised Bacterial Expression of a Synthetic BRIL Antibody. Acta Crystallographica Section F.
C Kirschbaum, JL Bennett, Q Tian, N Sen, IPS Smith, D Wu, JLP Benesch, S Khalid, GL Isom, CV Robinson (2025). Following phospholipid transfer through the OmpF3–MlaA–MlaC lipid shuttle with native mass spectrometry. PNAS.
BF Cooper*, R Clark*, A Kudhail, D Dunn, Q Tian, G Bhabha, DC Ekiert, S Khalid, GL Isom# (2025). Phospholipid transport to the bacterial outer membrane through the envelope-spanning bridge YhdP. Journal of Molecular Biology.
C Vieni, N Coudray, GL Isom, G Bhabha, DC Ekiert (2022). Role of Ring6 in the Function of the E. coli MCE Protein LetB. J Mol Biol
N Coudray*, GL Isom*, MR MacRae*, MN Saiduddin, G Bhabha and DC Ekiert (2020). Structure of bacterial phospholipid transporter MlaFEDB with substrate bound. eLife
LR Kolich*, Y Chang*, N Coudray*, SI Giacometti, MR MacRae, GL Isom, EM Teran, G Bhabha and DC Ekiert (2020). Structure of MlaFB uncovers novel mechanisms of ABC transporter regulation. eLife. PMID: 32602838
GL Isom*, N Coudray*, MR MacRae, CT McManus, DC Ekiert, G Bhabha (2020). LetB structure reveals a tunnel for lipid transport across the bacterial envelope. Cell. PMID: 32359438
GW Hughes, SC Hall, CS Laxton, P Sridhar, AH Mahadi, C Hatton, TJ Piggot, PJ Wotherspoon, AC Leney, DG Ward, M Jamshad, V Spana, IT Cadby, C Harding, GL Isom, JA Bryant, RJ Parr, Y Yakub, M Jeeves, D Huber, IR Henderson, LA Clifton, AL Lovering and TJ Knowles (2019). Evidence for phospholipid export from the bacterial inner membrane by the Mla ABC transport system. Nature Microbiology. PMID: 31235958
GL Isom, JL Rooke, CA Antunes, E Sheehan, TJ Wells, C Icke, AF Cunningham, JA Cole, IR Henderson and AE Rossiter (2018). Mammalian Cell Entry domains are required for bile resistance and virulence in Salmonella. Biorxiv
CA Antunes, EJ Richardson, J Quick, P Fuentes-utrilla, GL Isom, EC Goodall, J Möller, PA Hoskisson, AL Mattos-Guaraldi, AF Cunningham, NJ Loman, V Sangal, A Burkovski and IR Henderson (2018). Complete closed genome sequence of nontoxigenic invasive Corynebacterium diptheriae bv. mitis strain ISS 3319. Genome Announcements. PMID: 29437107
FC Morris, TJ Wells, JA Bryant, AE Shager, YR Sevastsyanovich, DJP Squire, J Marshall, GL Isom, J Rooke, R Maderbocus, TJ Knowles, M Overduin, AE Rossiter, AF Cunningham and IR Henderson (2018). YraP Contributes to Cell Envelope Integrity and Virulence of Salmonella enterica Serovar Typhimurium. Infection and Immunity. PMID: 30201701
GL Isom, NJ Davies, ZS Chong, JA Bryant, M Jamshad, M Sharif, AF Cunningham, TJ Knowles, SS Chng, JA Cole and IR Henderson (2017). MCE domain proteins: conserved inner membrane lipid-binding proteins required for outer membrane homeostasis. Scientific reports
DC Ekiert, G Bhabha, GL Isom, G Greenan, S Ovchinnikov, IR Henderson, JS Cox and RD Vale (2017). Architectures of lipid transport systems for the bacterial outer membrane. Cell. PMID: 28388411
